Growth & Wood Quality Breeding Tools
White spruce was our cornerstone species for molecular marker development and marker-assisted tree breeding applications. We conducted large-scale association studies in unstructured populations by genotyping thousands of SNPs and candidate genes (total of 20M genotypes). Our results suggested that the association genetics approach based on candidate genes would yield numerous markers to carry forward into validation and value demonstration experiments. Single-gene and multiple-gene approaches (MGS, multiple-gene selection) relying on Bayesian statistics were used to identify high priority candidate genes in which SNP coverage was intensified. We also tested and developed genomic selection (GS) by genotyping several hundred biparental progenies from breeding groups in the Quebec breeding population for thousands of gene loci.
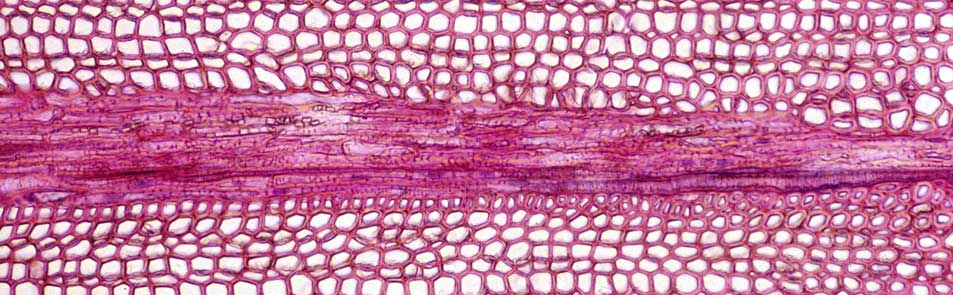
We aimed to validate diagnostic markers from previous and current association genetics work (see Figure). For example, breeding values from a large-scale first-generation breeding population were compared with predictions from MGS. We also showed that SNPs in expressed genes may be involved in adaptive population differentiation; therefore, we investigated diagnostic markers in natural populations with the aim of monitoring functional natural genetic diversity. These marker discovery approaches and findings were transferred to black spruce to broaden impacts and applications of this work in boreal forests. The outcomes of these objectives has been integrated to define marker systems that will assist tree breeding in two of the most important conifers and most reforested species for eastern and central Canada.
Goals
- Identify suites of diagnostic markers for variation in wood, growth and adaptation in juvenile and mature trees of white spruce, aiming to explain a useful part of the phenotypic variation for application in tree breeding.
- Support findings of marker-trait associations with functional and expression data.
- Evaluate the potential for using CNVs or PAVs as a source of sequence polymorphisms relevant for marker development.
- Validate groups of markers by testing marker/trait associations in independent populations and assessing the utility for MAS and GS in breeding populations developed by partners.
- Demonstrate the benefits of marker-based selections by measurement of wood yield and timber value recovery.
- Delineate range-wide patterns of diagnostic markers in natural populations of white spruce.
- Transfer to black spruce: identify and validate SNP markers of candidate genes for growth and adaptation.
Development of MAS for wood properties and growth in white and black spruces